CHD2 and CHD8
Personalized Modeling and Functional Analysis of ASD-related CHD2 and CHD8 Missense Variants
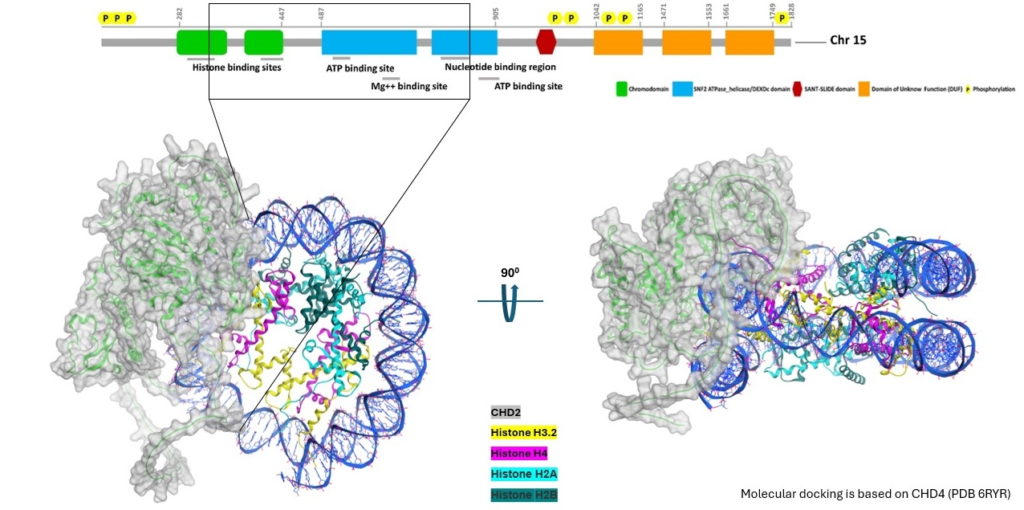
CHD2 (green) bound to nucleosome complex consisting of DNA (cyan), H4 (magenta), H3 (black), H2A (yellow) and H2B (blue). The red residues in the CHD2 protein are missense mutations of interest that we are investigating for their structural and functional impact on the CHD2 protein.
The chromatin remodelers, chromodomain-helicase-DNA-binding (CHD) proteins are among the most important epigenetic regulators, and are involved in biological processes such as cell stemness, proliferation, cell fate regulation, organ system development, and disease states such as cancer, and neurodevelopmental disorder. CHD2 and CHD8 proteins are important epigenetic regulators, and mutations in these are associated with neurodevelopmental disorders such as autism spectrum disorders (ASD) and intellectual disability (ID). Although next-generation sequencing (NGS) has considerably advanced clinical diagnostics, it has also pointed to major challenges such as the interpretation of variants of unknown significance (VUS), especially missense variants, where one amino acid is substituted for another. As the use of diagnostic sequencing grows in both clinical and research settings, the detection of VUS has become more common, particularly in the context of neurodevelopmental disorders. We are proposing to make personalized pathogenic missense and VUS isogenic cellular models of ASD using our own modified CRISPR/Cas9 strategy. CRISPR/Cas9 gene editing has played an important role in molecular biology over the past decade. We have already successfully generated several homozygous CHD2 missense cell lines this way. We aim to perform functional analysis on a set of 5 CHD2 and 5 CHD8 missense variants to understand the etiopathological mechanisms involved, such as protein function, stability/degradation, localization, and interactions with other proteins, compared to the wild type. Finally, we will perform computational analysis to identify compounds as molecular chaperones to stabilize and/or recover the expression and function of the mutated proteins. The ability to efficiently create disease models and correct risk variants offers significant potential for understanding the underlying mechanisms and therapeutic development.