NSUN2
In 2012, we reported in the American Journal of Human Genetics, back-to-back with the Human Molecular Genetics group at the Max-Planck Institute in Berlin, the discovery of NSUN2 as a gene for autosomal recessive intellectual disability (Khan et al, 2012; Abbasi-Moheb et al, 2012)).
The full article can be found at: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3376419/.
NSUN2 is also identified as the autosomal recessive intellectual disability locus MRT5 (http://omim.org/entry/611091).
Snap shots from the Khan et al 2012 paper are shown below.
Figure 1
A. Pedigree of family MR14 from Khairpur district. Filled circles indicate affected girls. B. HomozygosityMapper analysis25 for microarray SNP data: Genome-wide. Significant regions of HBD are seen only on 5p and 14q. The 14q locus was excluded because one of the unaffected siblings was also homozygous at this locus, whereas at the 5p locus, unaffected the sibling was genotyped as heterozygous. C. Ideogrammatic representation of the critical autozygous or HBD locus on 5p15.31, as determined from this study and in relation to the MRT5 locus identified by Najmabadi et al.8. D. c.2035G>A substitution encoding the Gly679Arg change in a heterozygous carrier and affected homozygote. E. ClustalW alignment of NSUN2 across multiple species showing conservation of the Gly679 residue in vertebrates and also in non-vertebrate animal species.
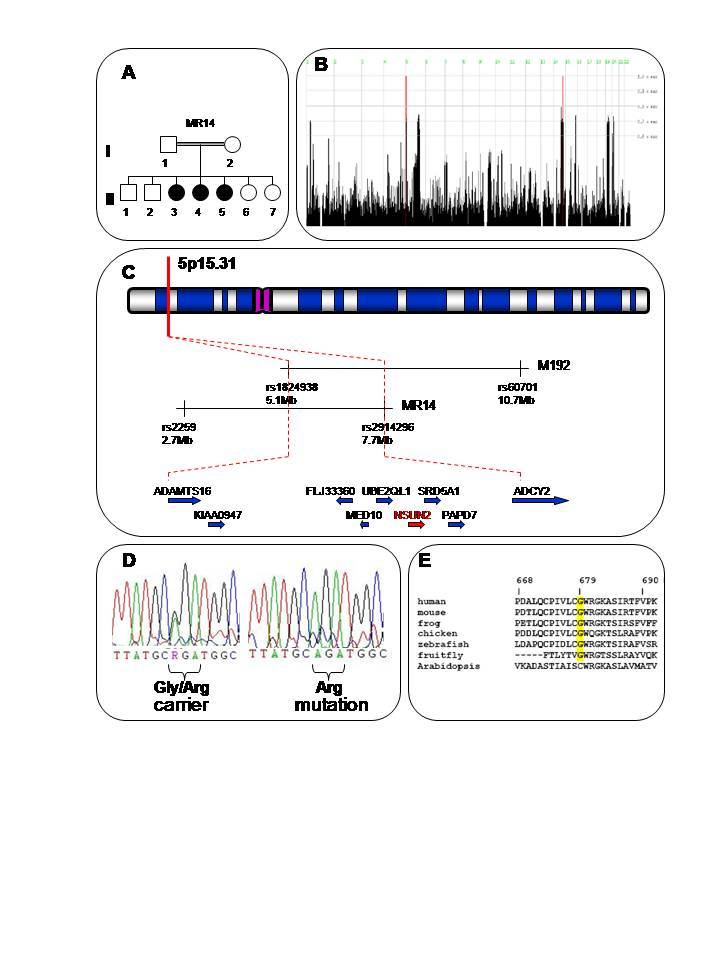
Figure 2
Wild type (WT) (A) and mutant (B) constructs for NSUN2 in the vector pcDNA-Myc transfected into the human breast cancer cell line HCC1954. A. WT NSUN2 (A) but not mutant NSUN2 (B) protein co-localizes with nucleophosmin, a nucleolar marker. (C) Co-staining for endogenous NSUN2 confirms exclusion of mutant NSUN2 (Myc-labelled) from the nucleoli. Arrows indicate nucleoli (A-C). Co-staining with DAPI shows nuclear localization.
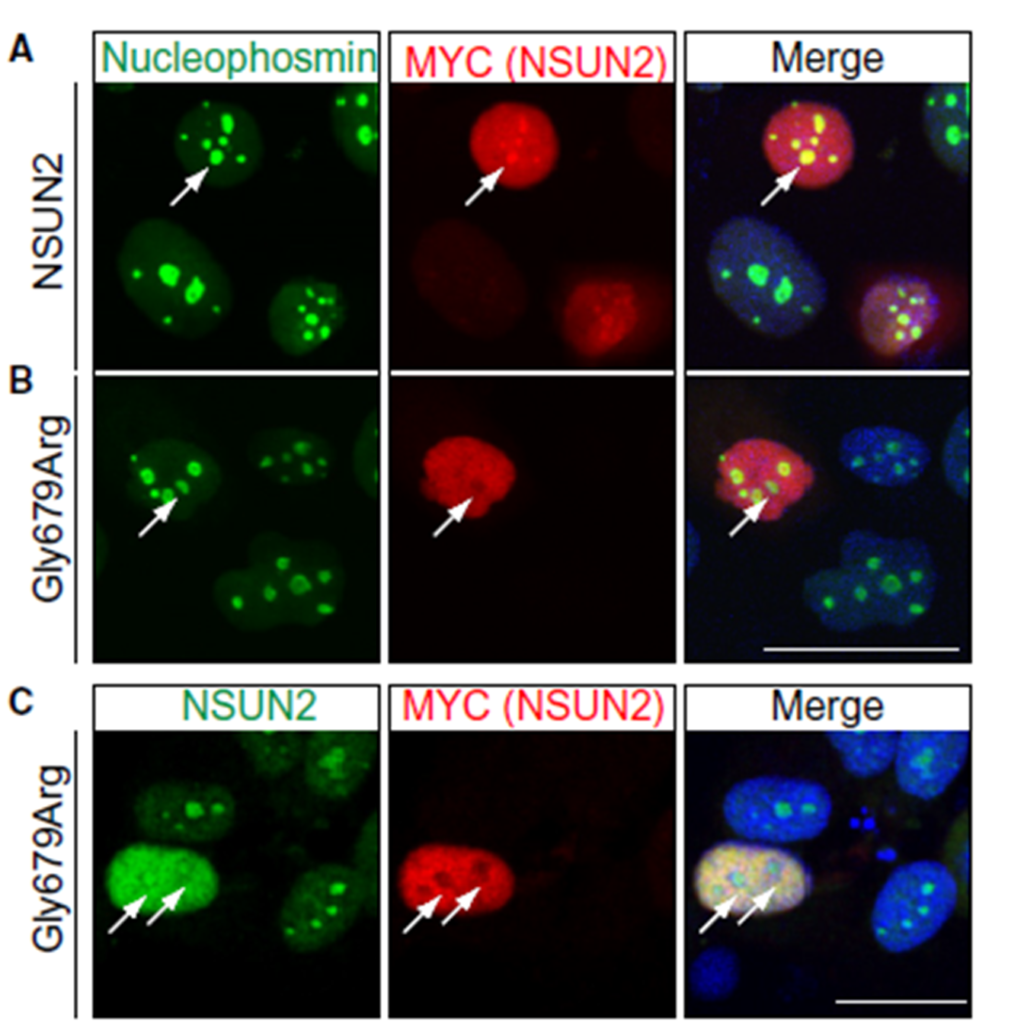
Figure 3
(A) Immunostaining showing nucleolar localization of wild-type (WT) NSUN2 in HCC1954 cells compared to cellular localization of NSUN2 carrying the 679Arg variant (Gly679Arg) in the nucleoplasm (B) and cytoplasm (C). Co-staining with DAPI shows nuclear localization. (D) Quantification of cellular localizations shown in (A-C). (E) Nucleolar localization of GFP-tagged wild type NSUN2 in HeLa cells versus (F) nuclear and cytoplasmic localization in 679Arg mutant NSUN2 (Gly679Arg). White arrow heads indicate nucleoli.
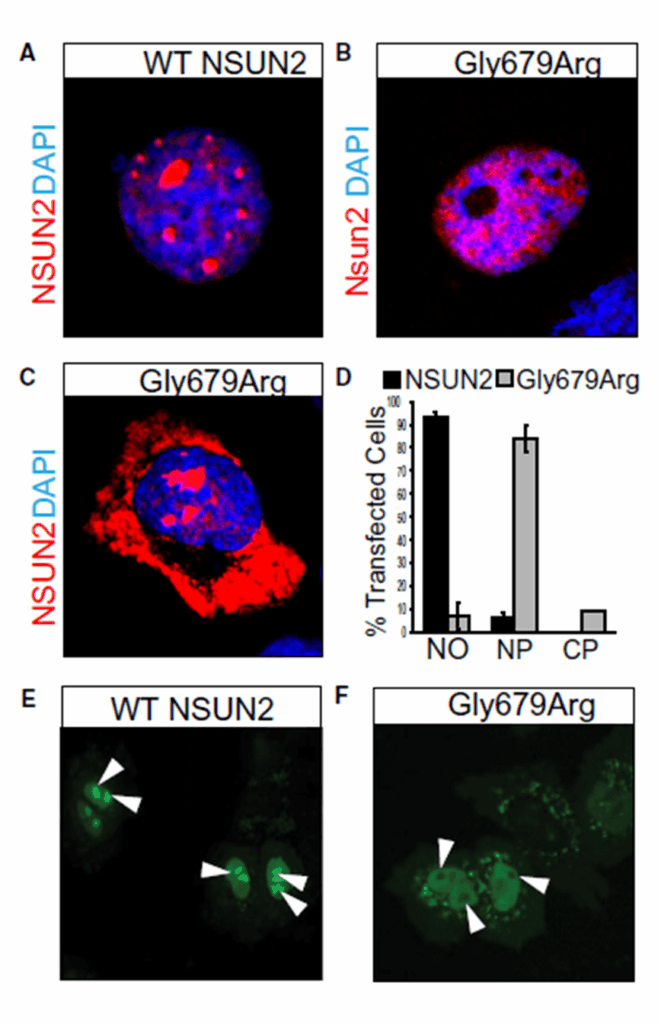
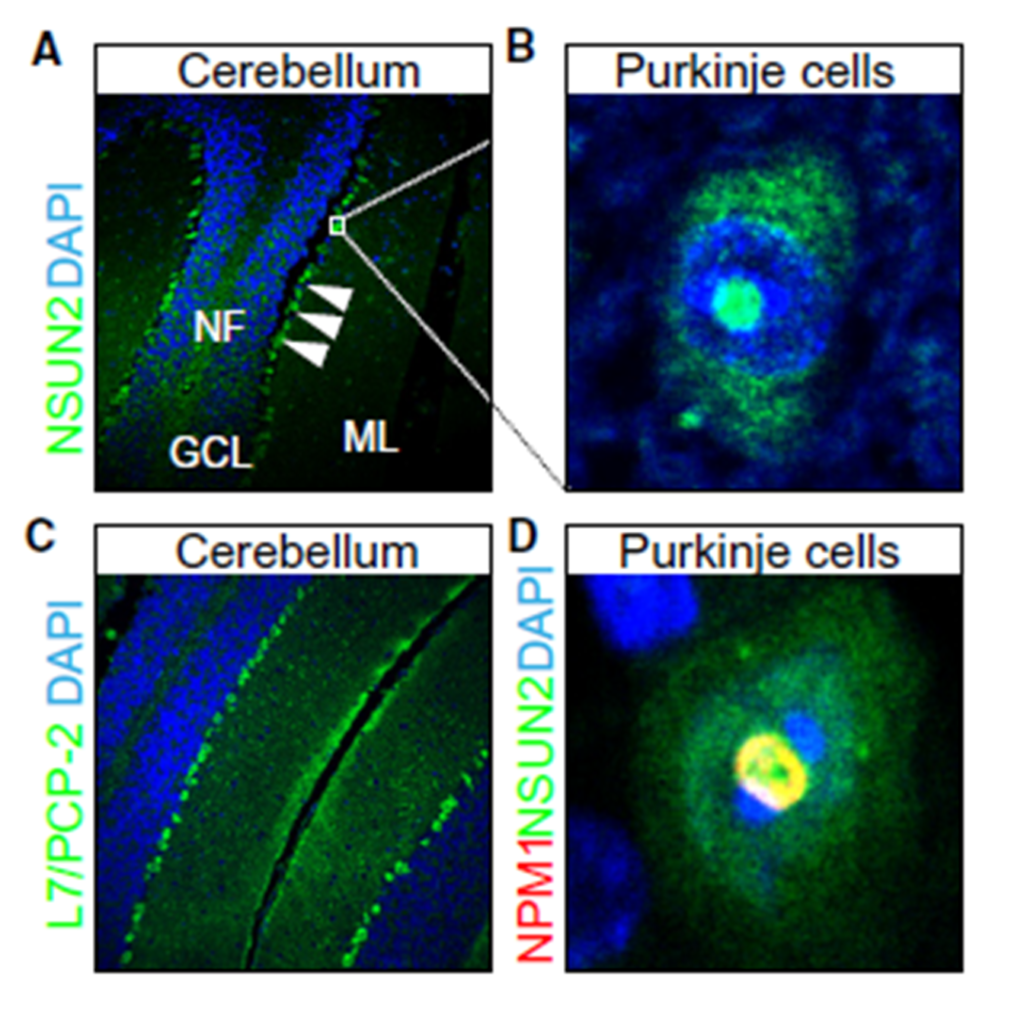
Figure 4
Sections of mouse cerebellum, (A) labeled with an antibody to NSUN2. Higher magnification of (A; insert) shows NSUN2 protein in Purkinje cells (B). NF: nerve fibers; GCL: granule cell layer; ML: molecular layer. (C) Sections of the cerebellum labeled for L7/Pcp-2, as a marker for Purkinje cells. (D) Co-localization of NSUN2 with nucleophosmin (Npm1) in nucleoli of Purkinje cells. Nuclei are counter stained with DAPI.